Mechanisms of Protein Stability Sensors: Structure and Function of Small Heat-Shock Proteins
Proteostasis in Health and Disease
Proteome fidelity, broadly defined as the preservation of the integrity of the structure and function of cellular proteins, is maintained through the coordinated action of many proteins, known collectively as the proteostasis network (PN). Members of the PN participate in protein synthesis, folding, disaggregation, or degradation and include the translational machinery, chaperones or co-chaperones, as well as the ubiquitin-proteosome system and the autophagy machinery. Failure of the PN has been associated with debilitating and fatal age-related diseases including cataract, Huntington’s disease, Alzheimer’s and Parkinson’s diseases.
Central to the PN are multiple families of chaperones or heat shock proteins (HSP) which supervise folding in the crowded environment of the cell, dynamically respond to changes in the environment that compromises the integrity of the proteome through rapid increase in the population of partially unfolding proteins, inhibit protein aggregation to the extent possible and participate in the clearance of aggregates.
Small Heat Shock Proteins
By possessing the largest capacity to bind unfolding proteins without the consumption of ATP energy, small heat shock protein (sHSP) are a first line of defense against aggregation. The sHSP/client protein complexes are a reservoir of refolding-competent clients for the HSP70 machinery after the conditions of stress abate.
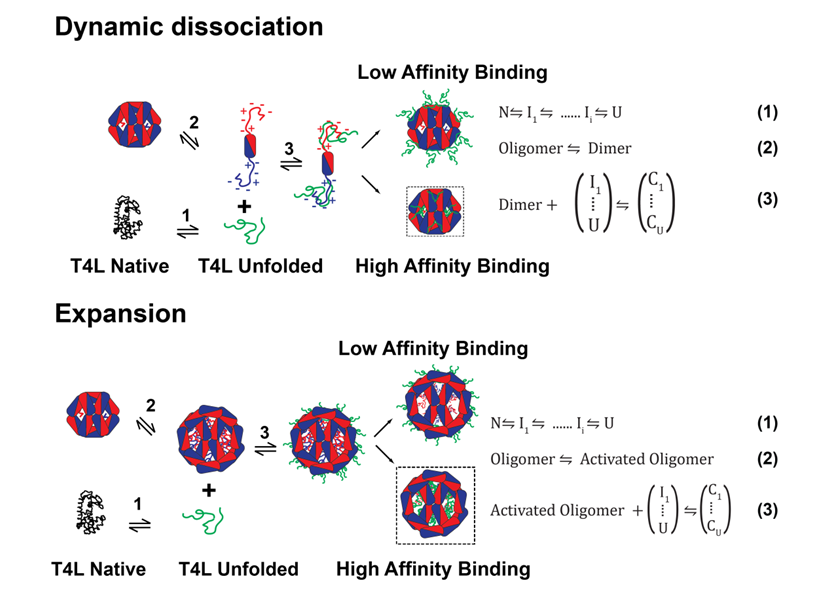
The Mchaourab Lab has developed has developed a general model for sHSPs chaperone activity that integrates structural and dynamic analysis of sHSP oligomers with thermodynamic evaluation of their interactions with client proteins . The resulting mechanistic principles, captured by the coupled equilibrium of equations 1-3.
sHSP detect changes in protein stability- whether induced by temperature, side-chain damage, or point mutations- prior to nucleation of aggregation. We demonstrated that sHSPs are stability sensors able to differentiate between mutants of the same substrate protein, T4 Lysozyme (T4L), that have identical structures but small differences in their ΔGunf. The affinities of human α-crystallins, Hsp27 [49], or the archeal Hsp16.5, correlate with the ΔGunf of the T4L mutants, a result captured by the coupling of equations 1 and 3. Our novel approach revealed that sHSPs bind their substrate in two modes distinguished by their affinity and number of binding sites. In the high-affinity binding mode, 4 sHSP subunits bind one substrate molecule while the stoichiometry is one to one in the low-affinity mode.
Far from being static “sponges” for unfolded proteins, sHSP oligomeric structures display a fascinating spectrum of conformational flexibility. In the last two funding periods, we demonstrated that polydispersity of mammalian sHSP oligomers is a mechanism for regulation of binding affinity and capacity. As illustrated in the Figure above, dynamic oligomer dissociation exposes a highly flexible N-terminal region containing putative substrate interaction sites which are otherwise inaccessible. Shifts in the equilibrium between large oligomers and dimers, e.g. by phosphorylation of Hsp27 and aB-crystallin, activate substrate binding. Alternatively, N-terminal region accessibility can be increased by oligomer expansion. Thus, the energetic threshold for substrate binding is also determined by accessibility of the sHSP binding regions, expressed in eq. 2 as an equilibrium between resting and active sHSP (sHSPa). A series of papers culminating with a high resolution structure of an expanded Hsp16.5 variant revealed that structural coupling between the N-terminal domain and the C-terminal tail conformation determines the size of the assembly. This model harmonizes existing data from multiple laboratories into a molecular mechanism of polydispersity, dissociation, expansion and activation.
Initially conceived with a mechanistic structural biology focus, this program has evolved to apply an iterative methodology that leverages advances in the genetics of zebrafish to test mechanistic models in the lens of a living organism and to tackle the questions of the cellular and physiological roles of sHSP in general (see the Zebrafish page). Owing to the central function of chaperones in aging, an in vivo mechanistic understanding of their function is not only of fundamental biomedical importance but also has a direct impact on translational efforts to deal with aging diseases involving protein aggregation.