Nociceptive neurons exhibit highly branched dendritic arrays
The sensation of pain is perceived by specialized sensory neurons or “nociceptors” that utilize elaborate webs of dendritic processes to detect harmful stimuli. These features of nociceptive neurons are universally observed and are thus likely to depend on conserved genetic programs. The mechanisms that govern the creation of these complex structures are poorly understood but recent studies in Drosophila have revealed that sensory neuron morphology is controlled by surprisingly large and diverse arrays of transcription factors (Moore et al. 2002; Grueber et al. 2003; Kim et al. 2006; Parrish et al. 2006).
PVD: A C. elegans Nociceptive Neuron
Most C. elegans neurons adopt a simple morphology with a single unbranched axon and dendrite (White et al., 1986). The PVD neurons are a striking exception to this rule. The PVDs are generated as a bilateral pair of neurons (PVDL and PVDR), each one traversing the length of the animal on either the left or right sides. Each PVD projects a single axon to the ventral nerve cord but displays a complex dendritic arbor that envelops the animal with extensive thin branches directly abutting the hypodermis or “skin” (Smith, Watson et al., 2010; Oren-Suissa et al., 2010). Genetic studies and calcium imaging experiments have established that the PVD neurons detect multiple classes of stimuli including high threshold mechanical force (“harsh touch”), hyperosmolarity and acute changes in temperature (Way and Chalfie, 1989; Chatzigeorgiou et al., 2010; Mohammadi et al., 2013). Thus, the PVDs have been proposed to function as nematode versions of polymodal pain-sensing neurons or “nociceptors.”
PVD adopts a highly ordered dendritic array
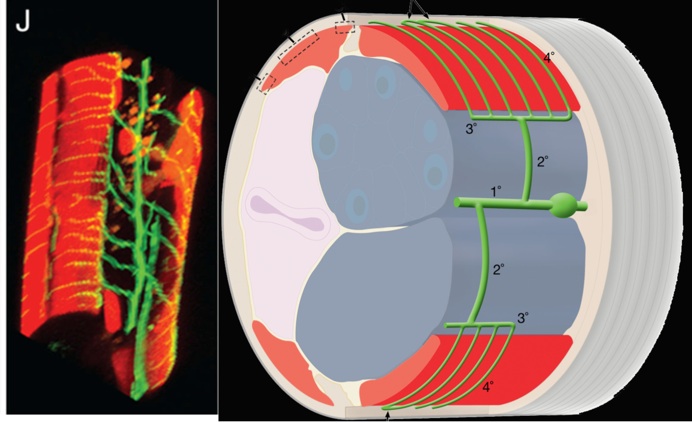
We used a bright cell-specific GFP reporter (Watson et al., 2008) to visualize PVD in the confocal microscope. Time-lapse imaging revealed that PVD dendrites adopt a series of orthogonal branching decisions to produce the elaborate adult structure (Fig. 2) (Smith, Watson et al, 2010) (Cover art) (for review see Shrestha & Greuber, 2010) . The close association of subsets of PVD branches with either longitudinal nerve cords or with motor neuron commissures suggests that these morphological landmarks may provide guidance cues.
UNC-6 (Netrin) drives dendritic self avoidance
We observed that the non-overlapping structure of the PVD arbor is correlated with the frequent occurrence of self-avoidance in which adjacent PVD dendritic processes withdraw upon contact with each other. The phenomenon of “dendritic self-avoidance” is widely observed but the mechanism is poorly understood (Corty et al., 2009). Our genetic analysis has revealed that the diffusible cue, UNC-6, and its receptors UNC-5 and UNC-40, are required to prevent PVD dendrites from invading the sensory fields of adjacent branches.

We have proposed a novel mechanism in which UNC-40/DCC captures UNC-6 at the tips of growing dendrites for physical contact with UNC-5 (Smith et al., 2012). These interactions drive dendrite retraction in a mechanism that depends on reorganization of the actin cytoskeleton. We have identified multiple downstream components of this pathway and are now investigating their roles in dendrite cytoskeletal dynamics.
Cell-specific microarray profiling reveals determinants of PVD dendritic morphogenesis
We used the mRNA tagging strategy to produce a gene expression profile of larval PVD neurons (Smith, Watson et al, 2010). This microarray experiment revealed > 2,000 transcripts that are enriched (> 1.5X) relative to all larval cells. We tested genes in this list with RNAi and genetic assays to reveal determinants of PVD branching. This approach detected eleven transcription factors that govern discrete steps in PVD morphogenesis. We are now extending the application of the mRNA tagging strategy to identify transcription factor targets with the idea that this approach should reveal suites of co-regulated genes that function together to specify PVD dendritic architecture.
Sensory neuron fates are distinguished by a transcriptional switch that regulates dendrite branch stabilization
With the goal of identifying new proteins with roles in dendrite branching, we used the mRNA tagging method to identify transcripts regulated by the LIM-homeodomain protein, MEC-3, (Smith, O’Brien et al., 2013) which is required for lateral branching in the PVD (Tsalik et al., 2003; Smith et al., 2010). A genetic and RNAi screen of candidate MEC-3 targets identified 17 hits with branching defects.
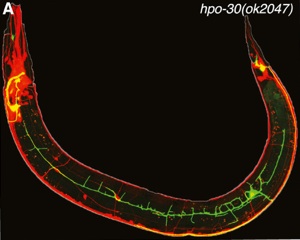
We confirmed that the claudin-like membrane protein, HPO-30, is regulated by MEC-3 and is required for stabilizing lateral PVD branches (Fig. 3). Because claudins are established components of intercellular junctions, we proposed that HPO-30 serves to link nascent PVD dendrites to the adjacent epidermis.
Additional genetic analysis revealed that MEC-3 and HPO-30 are regulated by a transcriptional switch involving the Aryl Hydrocarbon Receptor homolog, AHR-1. We showed that low levels of MEC-3 are sufficient to drive expression of hpo-30 in PVD but that high levels of MEC-3 are required to specify touch neuron fate. Our results show that AHR-1 provides the a dual function in touch neurons of activating MEC-3 expression while simultaneously repressing hpo-30 and other genes that contribute to PVD morphology and function. The phenotype of an ahr-1 mutant provides striking evidence of this role: The AVM touch neuron adopts the highly branched architecture and nociceptive sensory modalities of PVD while ectopically expressing hpo-30; mec-3 levels are reduced. Remarkably, the Drosophila ARH receptor homolog, Spineless, also regulates sensory neuron branching (Kim et al., 2006) which thus suggests that this role may be evolutionarily ancient and likely conserved in mammals (Smith, O’Brien et al., 2013)