Lab summary
Long noncoding RNA biology
Long noncoding RNA biology in mESC
Long noncoding RNA biology in mESC
Signaling in physiological environments in human cells
A rate threshold mechanism regulates MAPK stress signaling and survival
Kinetics of osmotic stress regulates a cell fate switch of cell survival
Predictive modeling and model-identification of signal transduction and gene regulation
Combining quantitative experiments with predictive modeling to understand cell signaling
Quantitative methods for single cell analysis
Overview:
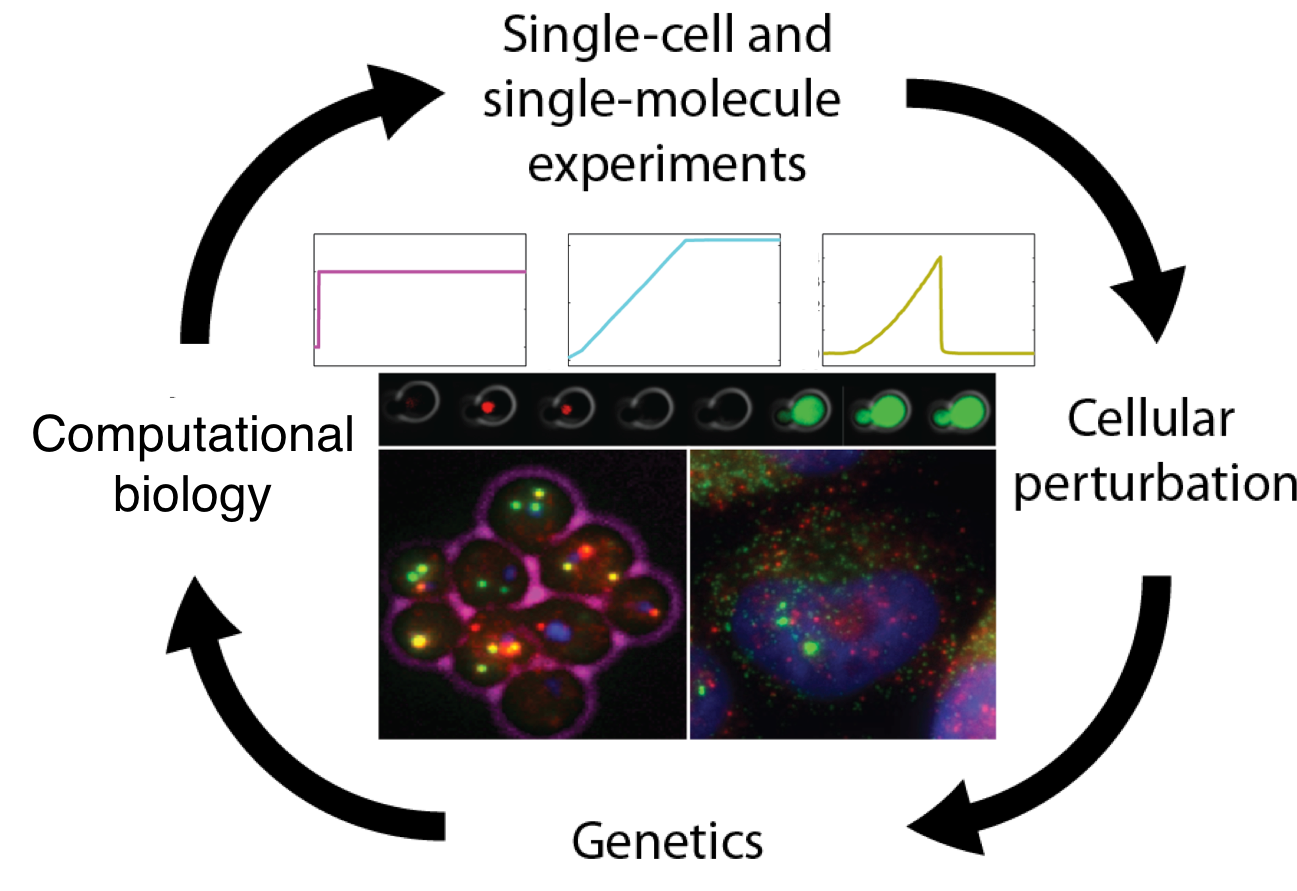
Numerous human maladies arise from alterations in cellular and molecular pathways stemming from exposure to pernicious environmental conditions. Alas, the lion’s share of biological investigations into the molecular changes that occur in diseased cells are carried out in environments that do not adequately replicate physiological conditions. Additionally, even individuals who are diagnosed with the same ailment may exhibit vastly dissimilar reactions to treatment, yet most studies assume that all cells of a given type behave uniformly. To surmount these limitations, the Neuert laboratory is dedicated to comprehending the fundamental mechanisms of signal transduction and gene expression in normal and diseased physiology within the context of kinetics and variability. A quantitative framework is employed to tackle broad biological inquiries, utilizing a diverse set of methods such as single-cell, single-molecule, and genome-wide assays, along with computational data analysis, genetics, molecular biology, chemical profiling, and single-cell predictive computational modeling. This integrative experimental and computational framework constitutes the bedrock of the Neuert lab’s expertise.